Welcome to the PGGB world!
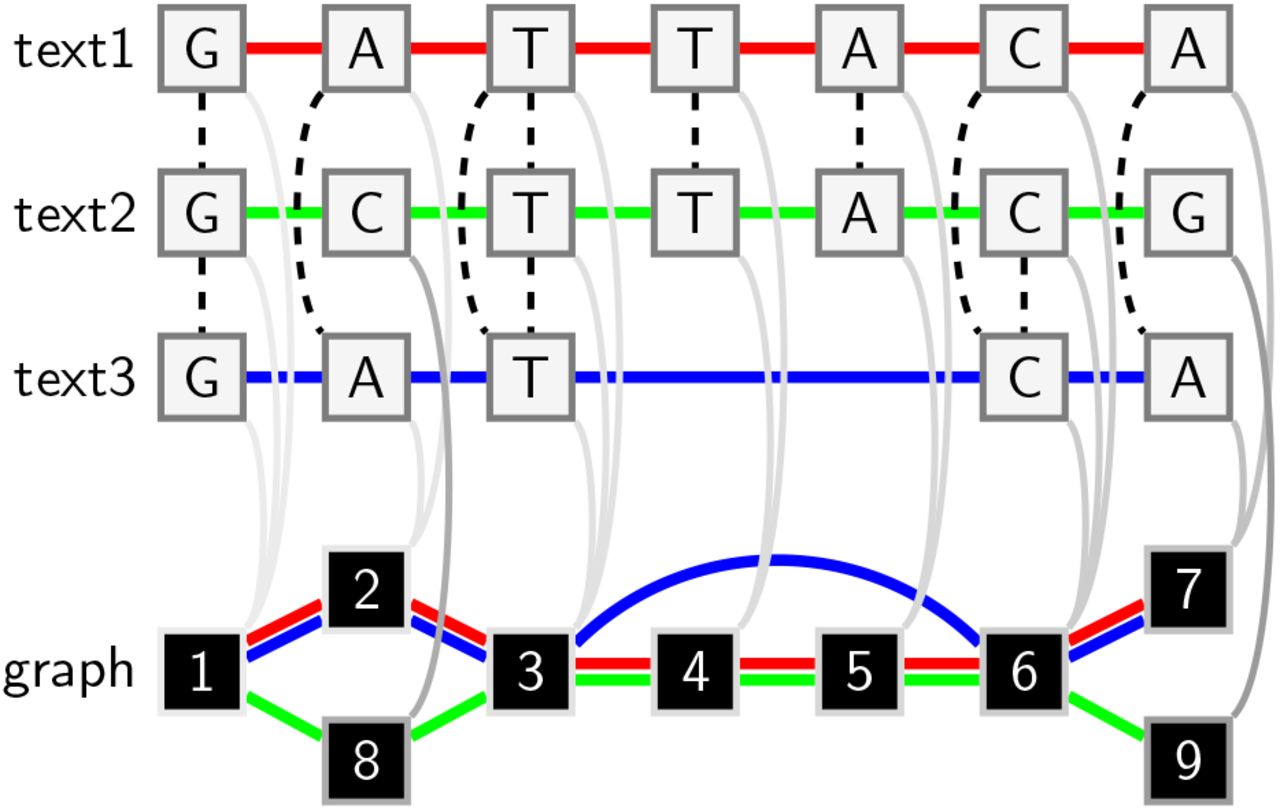
In standard genomic approaches sequences are related to a single linear reference genome introducing reference bias. Pangenome graphs encoded in the variation graph data model describe the all versus all alignment of many sequences.
pggb renders a collection of sequences into a pangenome graph, in the variation graph model. Its goal is to build a graph that is locally directed and acyclic while preserving large-scale variation. Maintaining local linearity is important for the interpretation, visualization, and reuse of pangenome variation graphs.
Core packages
Pairwise sequence alignment with wfmash
|
|
Graph induction with seqwish
|
|
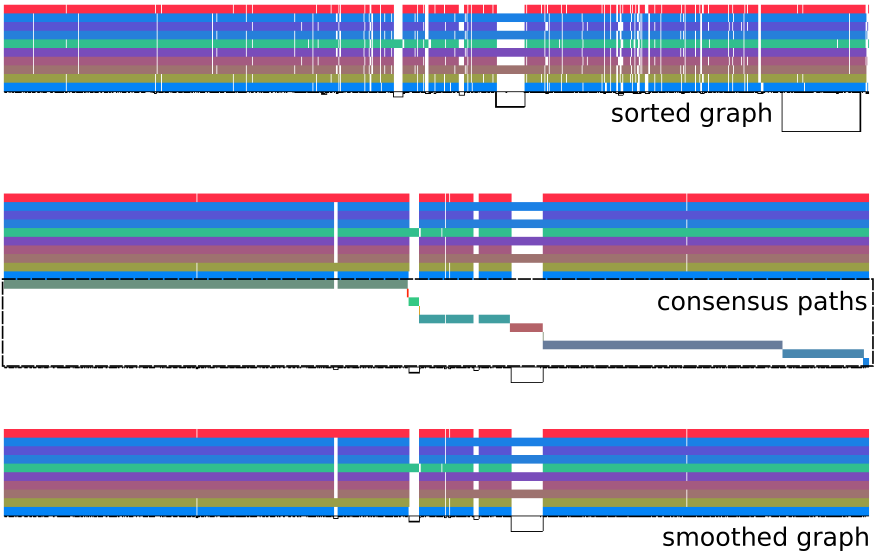
Graph normalization with smoothxg |
Contributed packages
Moreover, the pipeline supports identification and collapse of redundant structure with GFAffix.
Optional post-processing steps with ODGI provide 1D and 2D diagnostic visualizations of the graph and basic graph metrics.
Variant calling is also possible with vg deconstruct to obtain a VCF file relative to any set of reference sequences used in the construction.
It utilizes a path jaccard concept to correctly localize variants in segmental duplications and variable number tandem repeats.
In the HPRC data, this greatly improved variant calling performance.
The output graph (*.smooth.fix.gfa) is suitable for read mapping in vg or with GraphAligner.
A Nextflow version of pggb is currently developed on nf-core/pangenome.
This pipeline presents an implementation that scales better on a cluster.
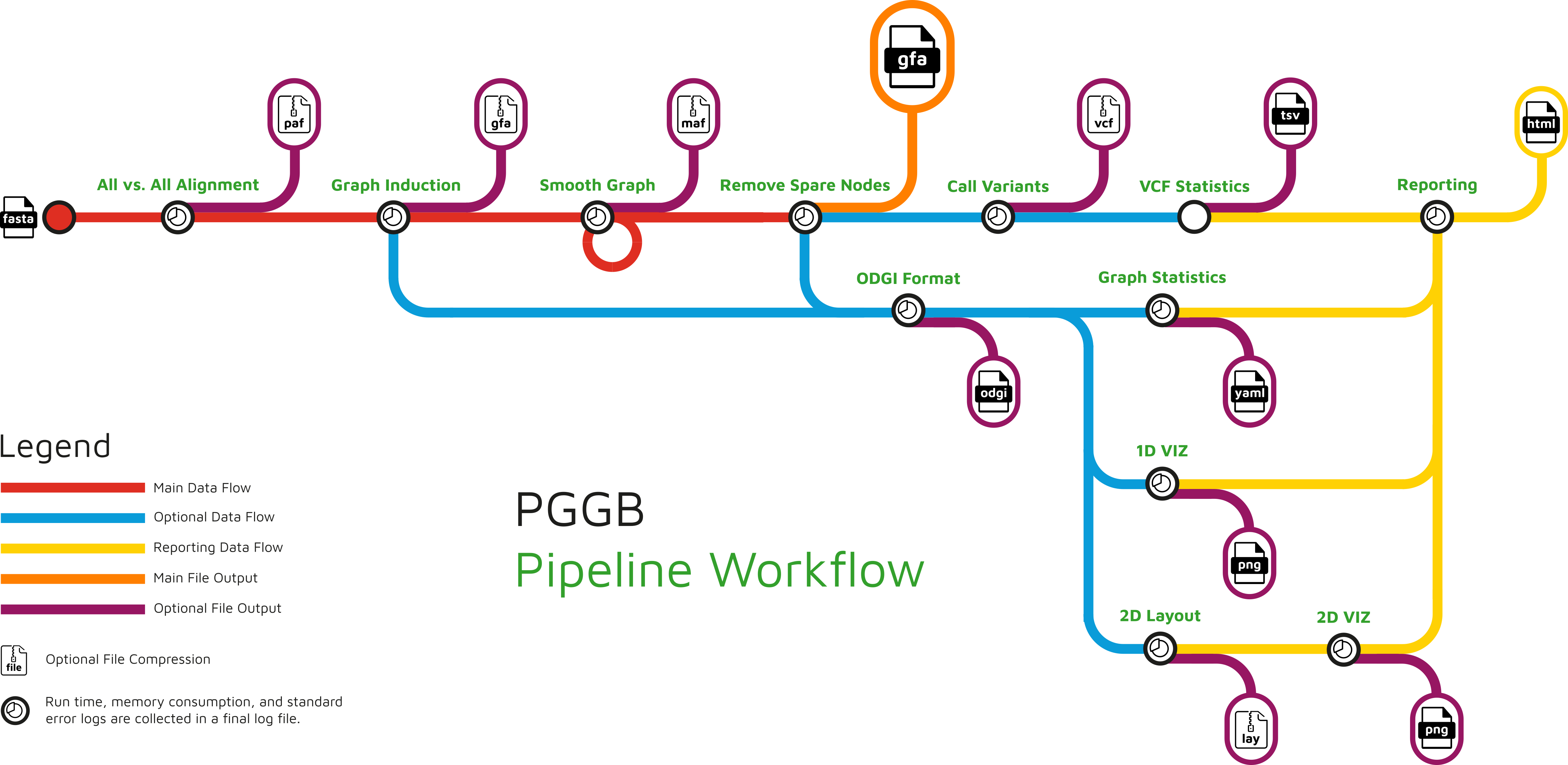
Pipeline Workflow